Reinforced Adversarial Neural Computer for de Novo Molecular Design | Journal of Chemical Information and Modeling
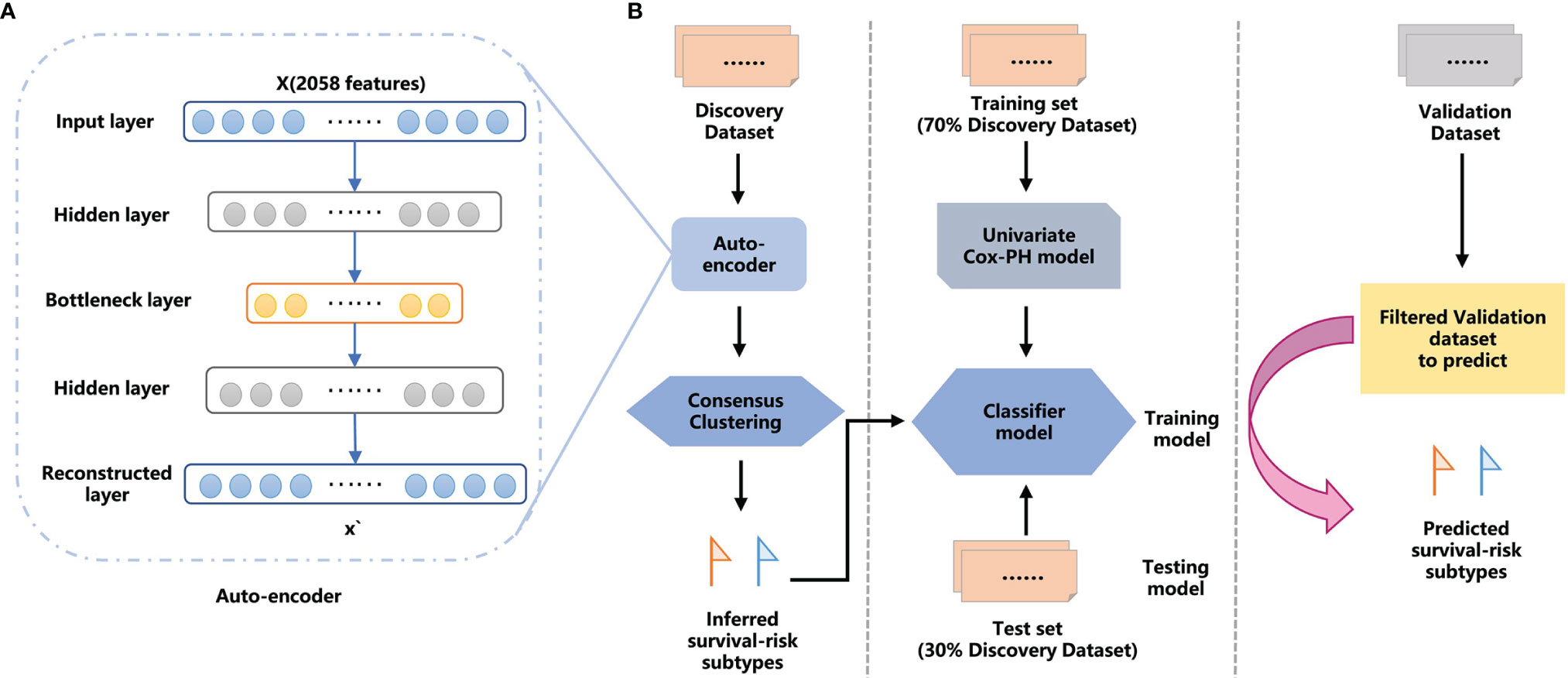
Frontiers | Deep Learning-Based Protein Features Predict Overall Survival and Chemotherapy Benefit in Gastric Cancer
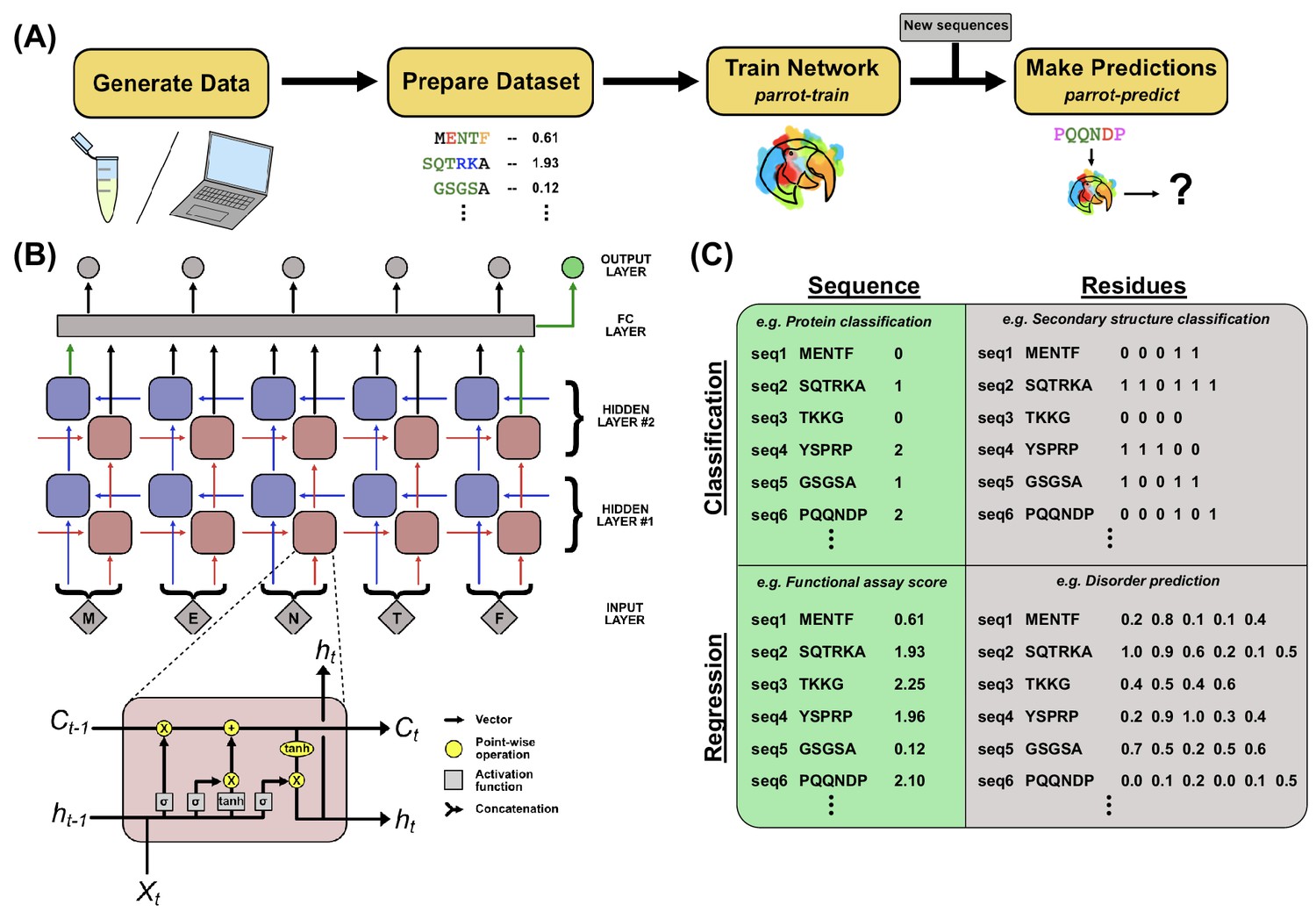
PARROT is a flexible recurrent neural network framework for analysis of large protein datasets | eLife

autoBioSeqpy: A Deep Learning Tool for the Classification of Biological Sequences | Journal of Chemical Information and Modeling
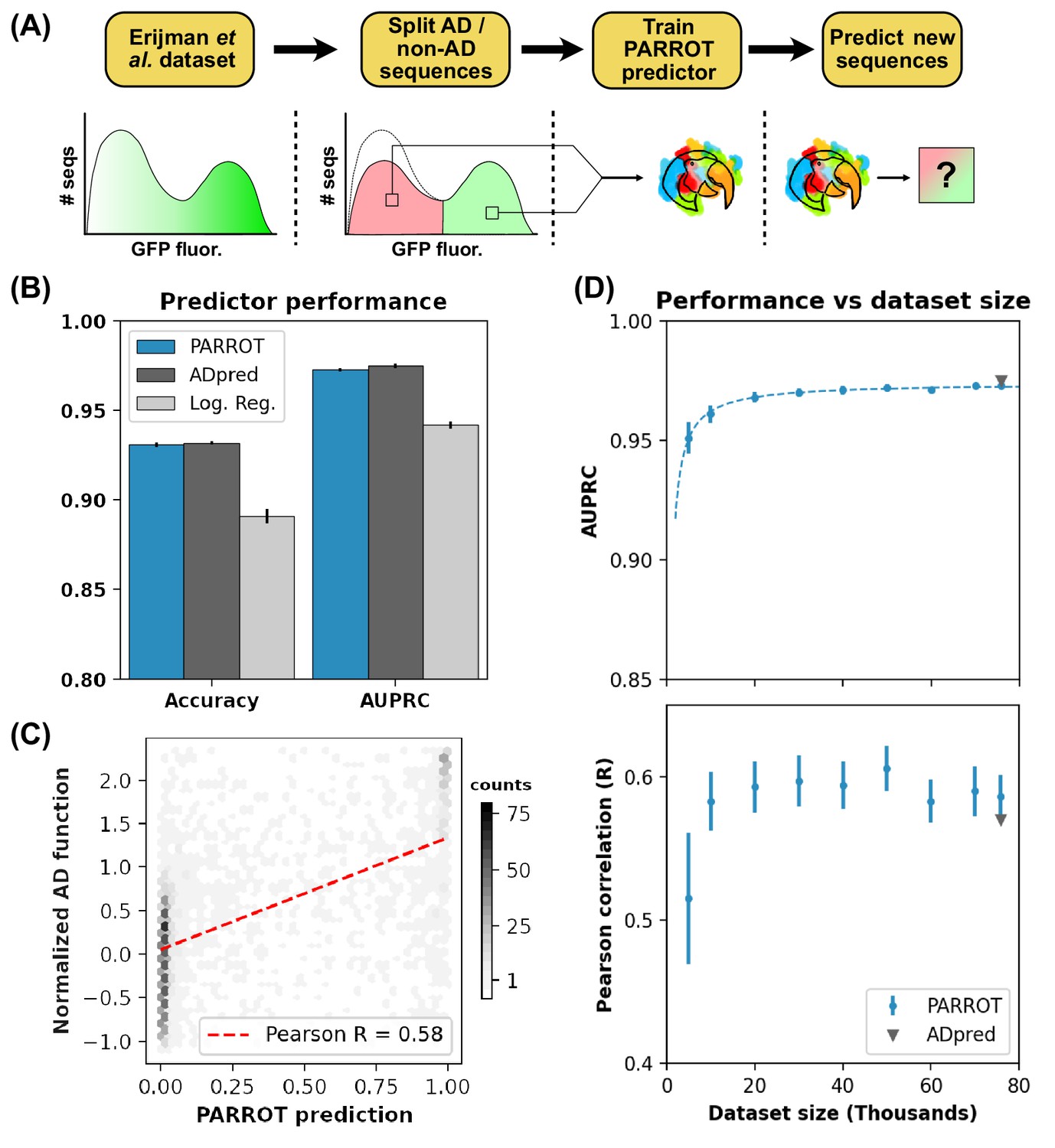
PARROT is a flexible recurrent neural network framework for analysis of large protein datasets | eLife
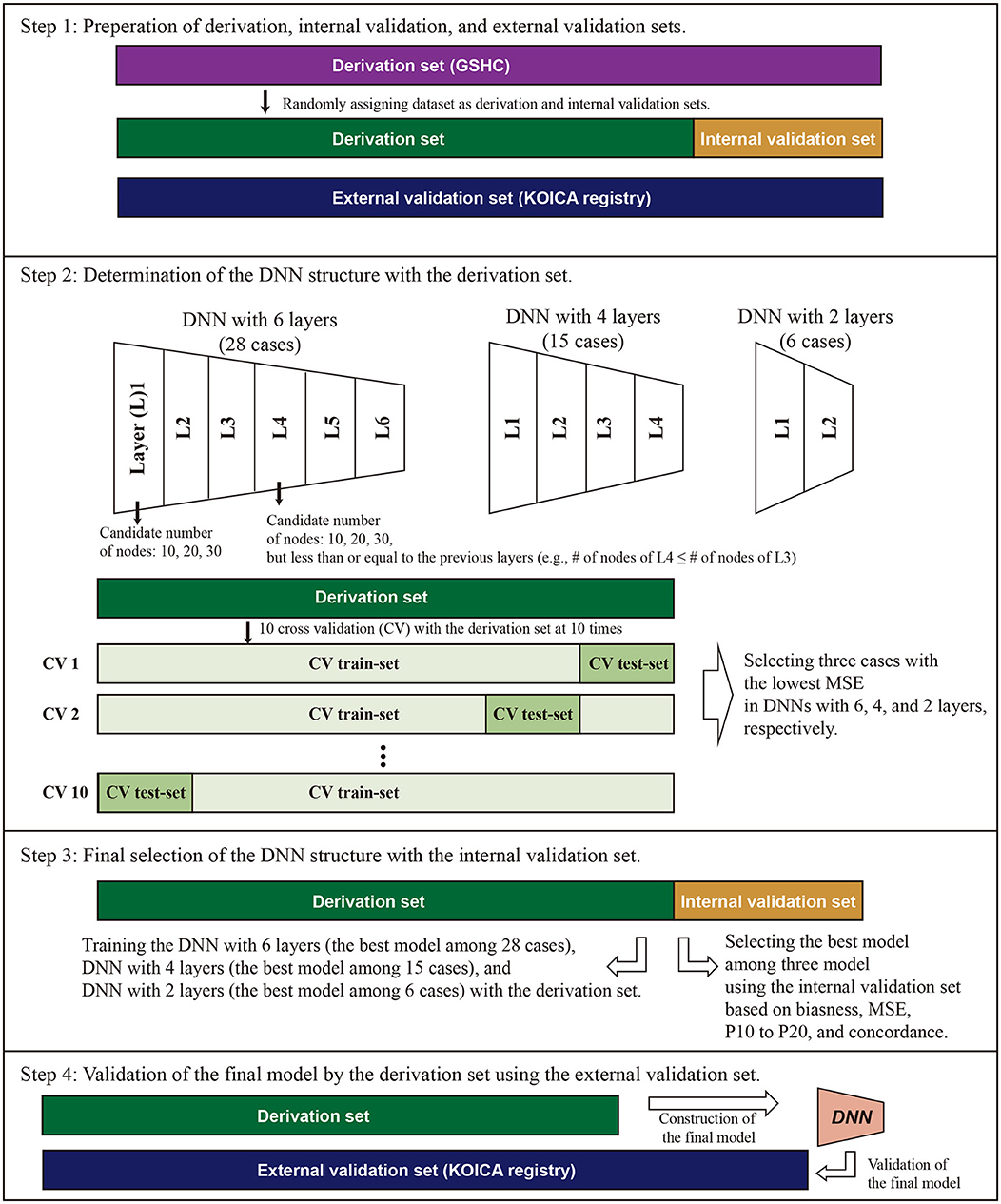

Frontiers | Comparison of a Machine Learning Method and Various Equations for Estimating Low-Density Lipoprotein Cholesterol in Korean Populations

Feature Extraction Approaches for Biological Sequences: A Comparative Study of Mathematical Models | bioRxiv

PDF) Critical assessment and performance improvement of plant-pathogen protein-protein interaction prediction methods
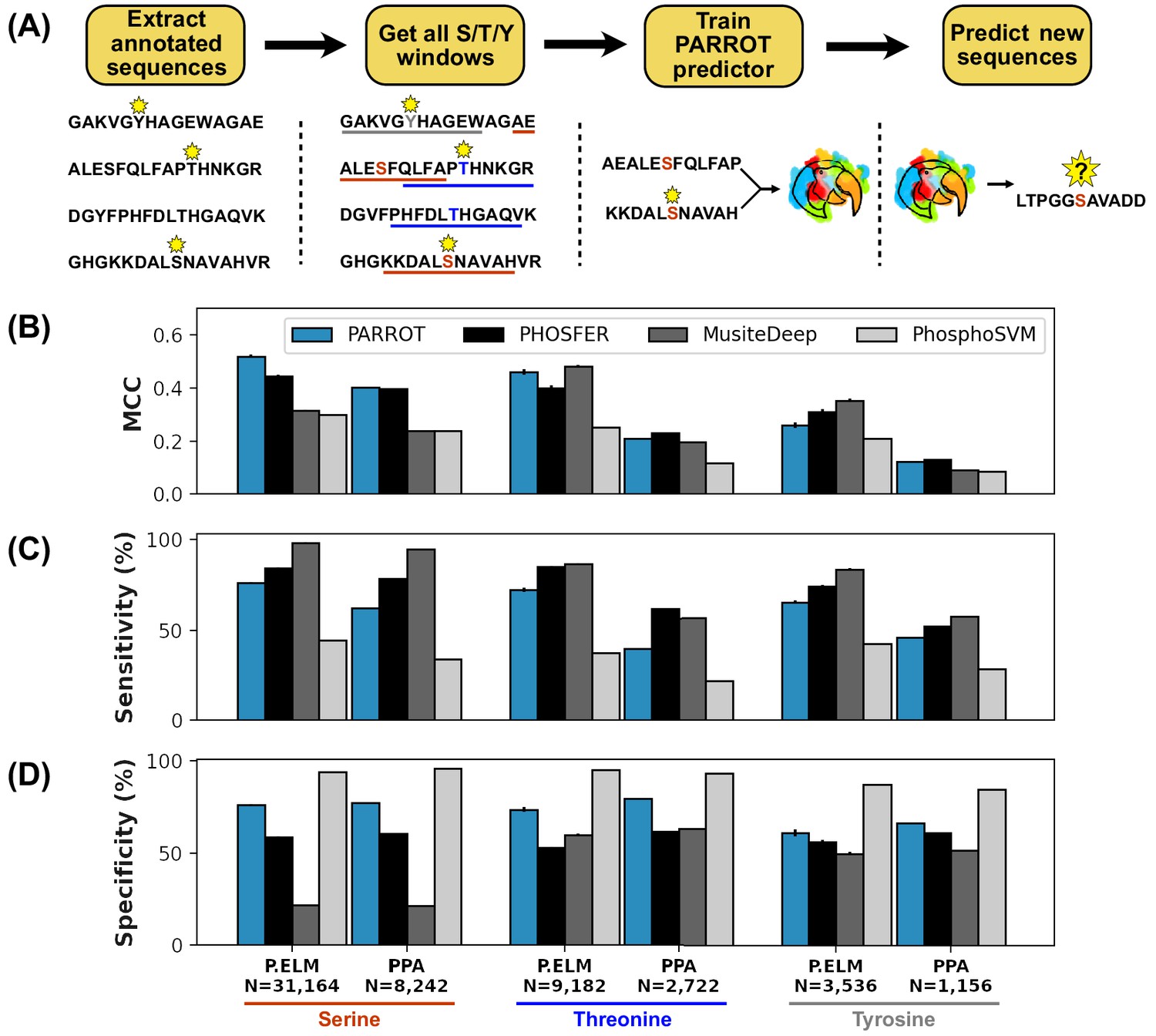
PARROT is a flexible recurrent neural network framework for analysis of large protein datasets | eLife

Improved sequence-based prediction of interaction sites in α-helical transmembrane proteins by deep learning - ScienceDirect
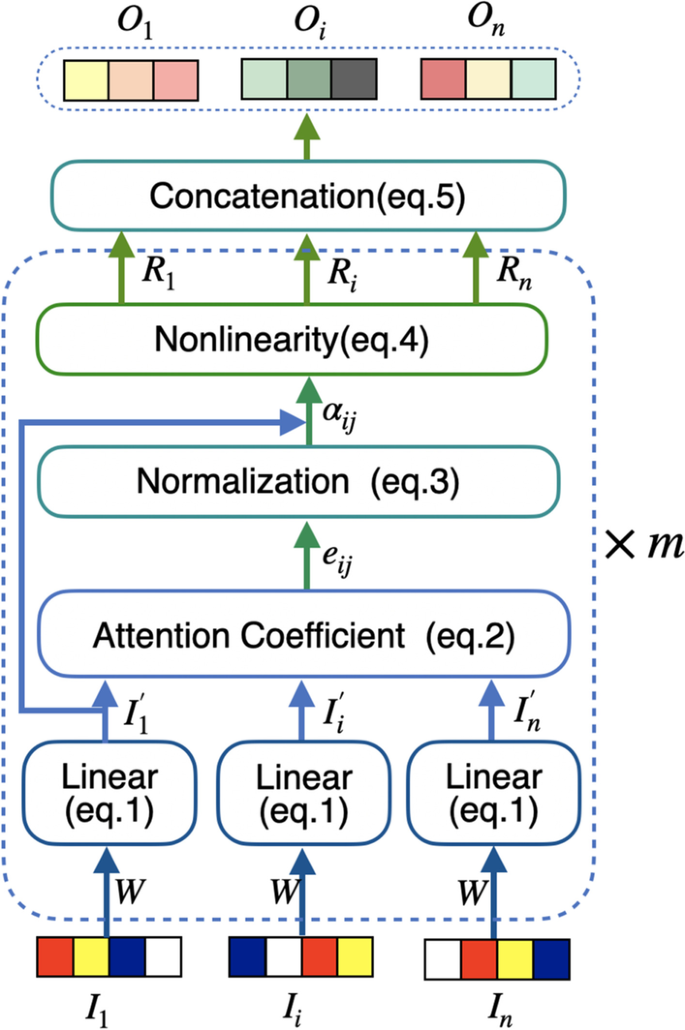
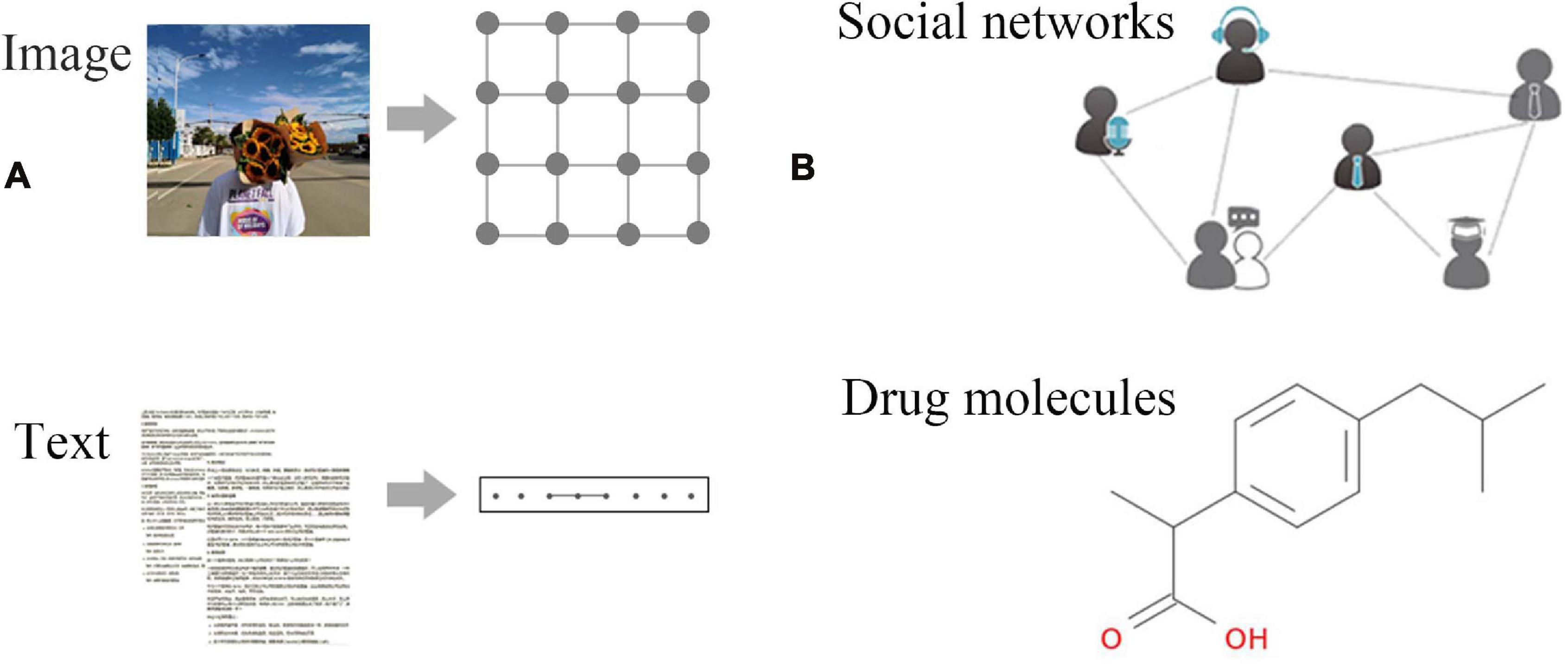
A novel hybrid framework for metabolic pathways prediction based on the graph attention network | BMC Bioinformatics | Full Text

Predicting Drug-Induced Liver Injury Using Convolutional Neural Network and Molecular Fingerprint-Embedded Features | ACS Omega